Strain*Treatment 1 0.13009 0.13009 0.13009 17.09 0.001Īs the strain*treatment p-value is 0.001, the null hypothesis that the strains are responding in the same way should be rejected at p<0.01, and we conclude that the strains differ in response.Īn interaction plot is given below which shows that the logWBC counts do not differ much in control mice of the two strains, but that following treatment with chloramphenicol they decline markedly in strain CBA but not in CD-1. General Linear Model: LogWBC versus strain, TreatmentĪnalysis of Variance for LogWBC, using Adjusted SS for Tests Below the table is an estimate of the pooled standard deviation S, and R-sq which indivates the proportion of the total variation accounted for by the two factors. These are because of the unequal numbers in each group. The table also has two columns “Seq SS” and “Adj SS”. As this is highly significant (p=0.001), this indicates that the strains do not respond in the same way (at the p=0.001 level of probability). This differs from previous tables in having a row called “strain*treatment” which indicates whether the strains are responding in the same way. The table first shows the factors (strain, treatment), their type (both fixed), their levels (2 in each case) and their values. The GLM-ANOVA output from on the logarithmically transformed data MINITAB is given below. However, one was added to each value before taking logs in order to avoid negative numbers (some numbers are less than one so the logarithm would have been negative). Accordingly it was decided to transform the scale, using a logarathmic transformation. Clearly the variation is not the same in each group (see Residuals versus Fitted Values where the variation is less in the left-hand four points than in the right hand points).
#Interaction plot minitab trial#
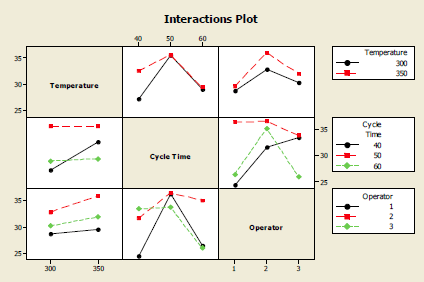
So in this case it is necessary to use a “general linear model analysis of variance” (GLM-ANOVA), which takes account of unequal numbers in each group.Ī trial GLM-ANOVA was done with the residuals plots (MINITAB 14) shown below.

Factorial designs are most easily analysed if there are equal numbers in each group, which is not the case here. White blood cell counts in controls and mice given chloramphenicol succinate at 2500 mg/kg Mice of each strain (of comparable age and from the same breeder) were assigned at random to each treatment group, dosed by gavage and after an appropriate time the mice were bled and the white blood cell counts were obtained.
These data were extracted, purely for purposes of illustration, from a larger study involving five strains and six dose levels. Table 1 shows WBC counts in mice of two strains kept as controls or treated with chloramphenicol. The aim of the study was to determine the effect of chloramphenicol on haematology of mice, and also whether strains differed in their response. The first is a 2×2 factorial showing what is meant by an interaction, and the second is a 4×2 factorial done using a randomised block design with two blocks.
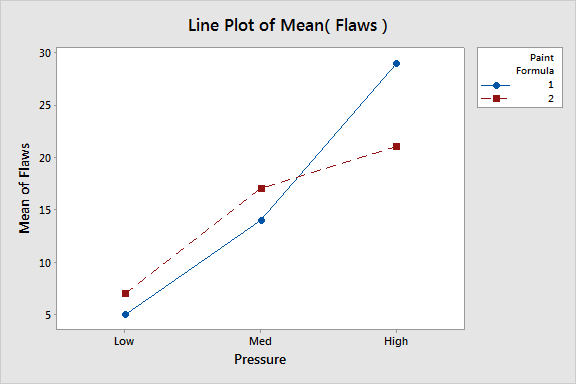
The purpose of factorial designs is to obtain more information from the same number of animals.
